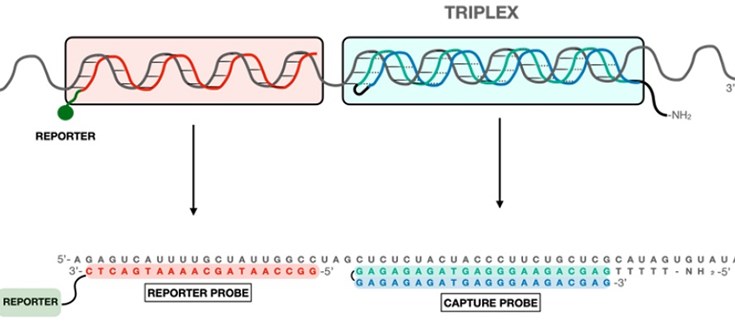
From the molecule to the bioassay by Custom antibody service (CAbS)-NANBIOSIS U2 as a PTI+Global Health Infraestructure
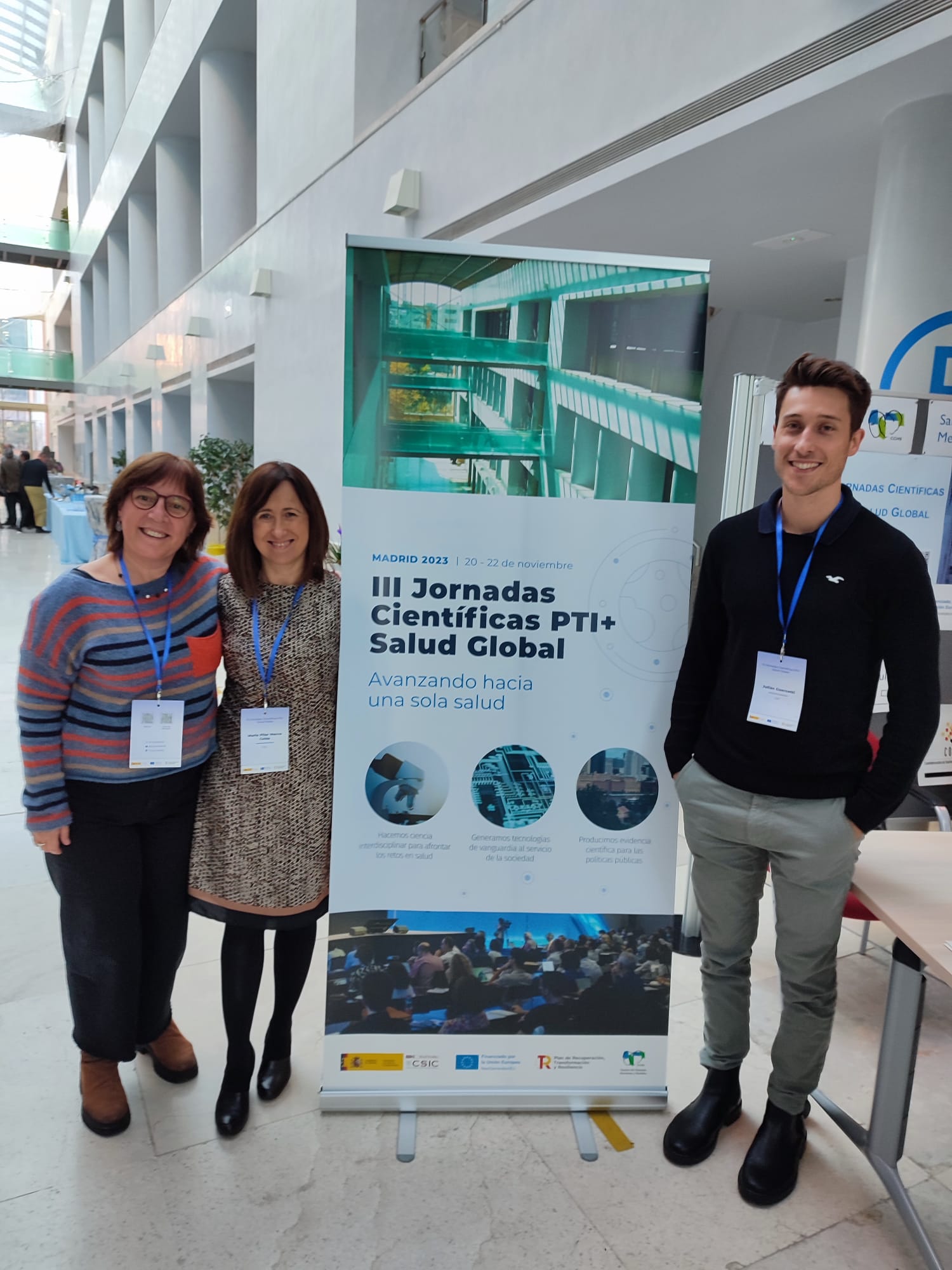
During 20-22 of November 2023, the III PTI+Global Health Scientific Conference were held in the Center for Human and Social Sciences, in Madrid.
In March 2020, the CSIC (Spanish National Research Council) launched the the Interdisciplinary Thematic Platform (PTI) on Global Health to bring together research teams and enhance knowledge about the new coronavirus SARS-CoV-2, which caused the pandemic. The PTI has mobilized and coordinates more than 400 scientists from 50 CSIC institutes in all areas.
The annual PTI+Global Health Scientific Conference are a meeting space where the results of the research carried out in the laboratories can be shown and discussed.
In the words of Margarita del Val, coordinator of the PTI+Global Health “In these III Conferences we are looking to the future to see how we evolve from the coronavirus to be prepared for future pandemics due to infectious diseases”. Iñaki Comas, coordinator of the PTI explained that this conference has been focused on “How to approach infectious diseases from a particular corner of knowledge but in an interdisciplinary way to be in a better position to face these global health challenges”.
The research caried out by the Nb4D group – NANBIOSIS U2 were presented by Julian Guercetti and Lluisa Villaplana:
“Towards a novel molecular signature for diagnosing infections based on Quorum sensing” M.-Pilar Marco; Juan Raya; Nuria Pascual; Nerea Castro; Carla Ferrero; J.-Pablo Salvador
“Immuno-μSARS2 chip: Correlating COVID-19 clinical severity with IgG personalized profiles” Julian Guercetti; Marc Alorda; Miriam Royo; Alicia Lacoma; Eduardo Padilla; Juan P. Horcajada; Silvia Castaneda; Agustín
Gutierrez-Galvez; Santiago Marco; J. Pablo Salvador; Pilar Marco, in this case with also with the participation of NANBIOSIS U3 Synthesis of Peptides Unit, led by Miriam Royo
“Using quorum sensing based antibodies as a new therapeutic strategy to treat Pseudomonas aeruginosa infections” Lluïsa Vilaplana Holgado; Bárbara Rodriguez Urretavizcaya; M.-Pilar Marco Colás
The Custom Antibody Service (CAbS) – NANBIOSIS U2 was presented by Julian Guercetti as one a PTI+Salud Global Infraestructure
“Custom antibody service (CAbS) from the molecule to the bioassay” Nuria Pascual Duran; Andrea Bastias; Idoia Camí; J.Pablo Salvador; Julian Guercetti; Lidia Hinojosa; Montserrat Rodriguez; Pilar Marco

Nanbiosis Unit 2 (Custom Antibody Service-CAbS) is a technological facility established in 2009 as part of the Spanish National Research Council (CSIC) and the Biomedical Research Center Network of Bioengineering, Biomaterials, and Nanomedicine (CIBER-BBN). Located within the Institute of Advanced Chemistry of Catalonia (CSIC) in Barcelona, the platform is equipped with a cell culture laboratory, housing the necessary equipment for obtaining, selecting, growing, and storing hybridomas. Additionally, the service offers laboratories for the synthesis of immunogens and the characterization of the produced antibodies.
The CAbS platform provides its monoclonal and polyclonal antibody production services to groups affiliated with CSIC and CIBER-BBN, as well as other research groups from public or private institutions and companies.
The primary goal of the service is to deliver high-quality service and scientific guidance in the production of immunoreagents, including polyclonal, monoclonal antibodies, and antibody fragments, as well as various probes such as protein and enzyme bioconjugates, biotinylated and fluorescent probes, biofunctionalized particles, and more.
The service is adaptable to each client’s needs and can produce antibodies against proteins, peptides, organic molecules, or other antigens through standardized or customized protocols. Special emphasis is placed on the immunogen design phase, a crucial aspect for modulating antibody selectivity and affinity.
One distinguishing feature of the CAbS service is its provision of guidance and assistance in preparing immunogens and producing antibodies for low-molecular-weight molecules, such as pigments, hormones, or anabolic agents. Service management is overseen by the NB4D group at IQAC-CSIC, a team with extensive experience in this field. Each service request is reviewed by a Scientific Committee, which produces a feasibility report before project acceptance. Users are kept informed of project progress at all stages and are consulted before proceeding based on the achieved results.
The services offered by the platform include:
• Preliminary discussion of project characteristics
• Design and synthesis of haptens
• Preparation of bioconjugates
• Hybridoma development
• Production of monoclonal antibodies
• Production of polyclonal antibodies
• Additional services (antibody purification, monoclonal antibody isotyping, etc.)
• Guidance and setup of immunoassays.
Recently, the unit has acquired a Surface Plasmon Resonance (SPR) instrument, which enables real-time detection and monitoring of interactions between two or more molecules without the need for labelling. The studies conducted with this instrument serve to determine specificity between compounds and/or characterize the kinetics and affinity of
these interactions. This SPR was funded by the European Commission – NextGenerationEU (Regulation EU 2020/2094), through CSIC’s Global Health Platform (PTI Salud Global).